How can the coronavirus mutate so quickly? It seems like every week there is a new variant that messes everything up. A new collaborative study from Oxford and Dundee could help us to understand exactly which mutations will be the most deadly, allowing us to design new vaccines before its too late.
When organisms (and viruses) reproduce, they must copy their genetic material so that the offspring has all the instructions needed to grow and survive. The parent’s double helical DNA is unzipped and each strand acts as a template for its own replication. Adenine must always pair with thymine, and guanine with cytosine.
This ingenious replication mechanism of DNA was first theorized by Watson and Crick, right after solving its 3D structure. At the end of their famous 1953 paper Francis offhandedly stated, “It has not escaped our notice that the specific pairing we have postulated immediately suggests a possible copying mechanism for the genetic material”.
The enzymes that copy your DNA are incredibly efficient but they sometimes make mistakes. Roughly 1 in 100 million Bases are paired incorrectly. This incredibly high accuracy is only possible due to a proofreading site inside the DNA polymerase complex: When the wrong base is added, a little kink is formed in the double helix and it gets lodged inside the enzyme. A magnesium ion will catalyze its removable so that the polymerase site can try again.
The enzymes that copy most viral genomes do not contain these proofreading sites, so far more errors slip through the cracks. HIV for example contains an RNA-dependant RNA polymerase enzyme that makes a mistake every 1 in 10000 base pairs!
Viruses give rise to thousands of offspring every single day so it doesn’t matter if most of them are rubbish. It’s a bit like the famous scenario of a thousand monkeys on typewriters trying to produce the complete works of Shakespeare. The few that randomly acquire an advantageous mutation will be even better at infecting cells and the cycle continues.

Humans are only capable of producing one baby every 40 weeks. If they have a damaging mutation, it is highly likely that they will die and no innovation will be passed on at all. This is why it takes millions of years for dramatic changes to appear in higher organisms, but viruses are reclassified by the week!
I’ll stop ignoring the elephant in the room now and begin to explain why this knowledge is so important to us at the moment.
Since its discovery in 2019 SARS-CoV-2 has rapidly spread around the world, gradually mutating to best suit the local environment and population. When this happens to a significant degree, epidemiologists will name it after the Greek alphabet so its development and spread can be tracked.
In December 2020, an Alpha variant was designated due to its 40-80% higher transmission rate. The most famous recent example is the delta variant. It was first sampled in India last year, then named in May 2021. As of October, there are >858,000 cases in the UK alone and it has spread to nearly 200 countries.
Mutations are often described in abstract terms and it is difficult to visualize exactly what is happening to the virus to make these new variants so infectious. In order to understand this we will take a look at the spike protein on a molecular level.
The surface of the coronavirus is covered in antigens called spike proteins. In order to infect a cell this spike must form a strong bond with a protein on our lung cells outer membrane called ACE2. This tricks the cell into letting the virus inside.
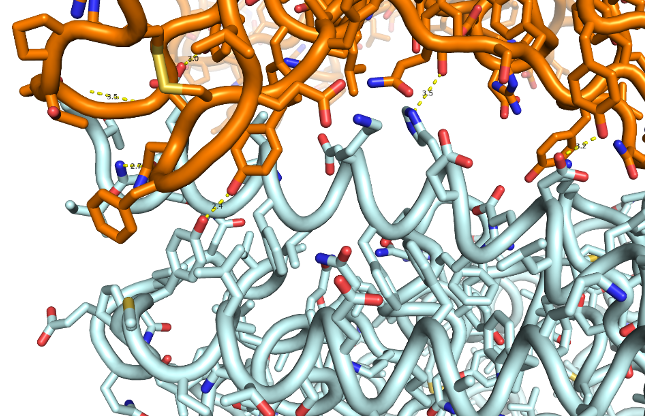
Proteins are made up of a long chain of amino acids, curled and folded into a specific 3D shape. When 2 proteins come together, a Velcro like bond is formed. Each amino acid contributes a weak electrostatic force of attraction but all together they are strong enough to hold the 2 surfaces together.
The key function of DNA (or RNA in the case of the coronavirus) is to code for the order of amino acids in a protein. If one of the DNA bases is changed, there is a chance that one amino acid will be substituted for one of the other 20 common choices.
Each amino acid has a different ‘functional group’ with varying chemical properties. Some hydrophobic, some hydrophilic. Some positively charged, some negatively charged, etc. The binding surface of ACE2 is overall negatively charged so if a neutral amino acid in the spike protein mutated to become positive: the bond would become stronger.
Alternatively, a mutation could occur higher up in the protein that makes it more evasive of the immune system. Several amino acids in spike accumulate bulky sugar side chains that shield the surface from proteins of the immune signals that recognize ‘pathogen associated molecular patterns’ (PAMPs)
The spike protein is over 1200 amino acids long so it if often difficult to identify whether mutations in a newly identified sample will make it more infectious or not. How can we measure this before thousands of people start to get infected?
In August 2021, researchers from Oxford and the University of Dundee made incredibly improvements to analytical techniques such as ‘surface plasmon resonance’ and ‘bilayer interferometry’ that made their measurements of Spikes binding affinity with ACE2 for more accurate.
The pitfalls of previous experiments were that this process was often measured at lower temperatures ~20’C to stop the proteins from denaturing. That meant that the kinetics would be far slower than reality. Many studies did not even report their temperature at all! This new experiment managed to stabilize it at a physiological temperature of 37’C.
Another common issue with older studies is that proteins tended to aggregate on the detector meaning that the Spike got tangled up and it wouldn’t dissociate as much as it should. This time, a far lower concentration was used and the sensitivity of the measurement was cranked up.
The key finding of this research is that mutation that provide the greatest infectiousness and dominance in their respective regions are in the receptor binding domain (RBD) of spike.
These findings correlate strongly with epidemiological studies for example: A mutation of asparagine-501 into a tyrosine amino acid is what helped the alpha variant to become so infectious. Kinetic analysis of this N501Y mutation proved that it binds to ACE2 five times more strongly.
Now that these affinity measuring techniques are more accurate to biological reality, scientists will be able to identify particularly deadly variants far more early. Communities can be quarantined and vaccinated before it spreads across the entire world. Thousands of lives will be saved.
If you would like to learn more about these experiments. Check out their original paper at Effects of common mutations in the SARS-CoV-2 Spike RBD and its ligand, the human ACE2 receptor on binding affinity and kinetics | eLife (elifesciences.org)
Featured image: A SARS-CoV-2 virus approaching a cell with ACE2 receptors.
Images: Matthew Clark


